Prevalence of Staphylococcus aureus in the Oral Microbiome of Children with Orofacial Clefts: A Comparative Study using PCR and Vitek Assays
| Received 19 Jan, 2026 |
Accepted 16 Apr, 2026 |
Published 30 Jun, 2026 |
Background and Objective: Cleft lip and palate (CLP) are among the most common congenital craniofacial anomalies, arising during early intrauterine development and affecting a substantial number of newborns worldwide. Individuals with oral clefts are prone to bacterial colonization, increasing the risk of infection and antibiotic resistance. The detection of methicillin-resistant Staphylococcus aureus is therefore clinically important. This study aimed to identify the presence of the mecA gene in S. aureus isolates obtained from patients with cleft lip and cleft palate and to evaluate the agreement between molecular and automated diagnostic methods. Materials and Methods: A laboratory-based analytical study was conducted on S. aureus isolates recovered from patients with oral clefts. Bacterial identification was performed using the Vitek automated system. Genomic DNA was extracted, and polymerase chain reaction was carried out to detect the mecA gene, followed by agarose gel electrophoresis. The diagnostic performance of PCR and Vitek was compared, and agreement between the two methods was assessed using the kappa coefficient. Data were analyzed in SPSS, with PCR-Vitek agreement assessed by Cohen’s kappa and cleft-S. aureus associations by ORs (95% CI; p<0.05). Results: The PCR analysis demonstrated distinct amplification bands corresponding to the mecA gene in several S. aureus isolates, confirming the presence of methicillin resistance. The Vitek system showed comparable identification results, with a notable level of agreement between the two diagnostic approaches. The findings indicate a considerable prevalence of mecA-positive S. aureus among individuals with cleft lip and palate, suggesting an increased risk of antibiotic-resistant infections in this population. Conclusion: The study highlights the relevance of combining molecular and automated diagnostic tools for accurate detection of methicillin-resistant S. aureus in patients with oral clefts. Early identification of resistance markers such as mecA can support effective infection control and guide appropriate antimicrobial therapy. Further studies with larger sample sizes are recommended to strengthen clinical correlations and resistance surveillance.
| Copyright © 2026 Hussein Abass et al. This is an open-access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. |
INTRODUCTION
Orofacial clefts, including cleft lip, palate, and both combined, are among the most common congenital anomalies worldwide, occurring in roughly 1 in 500 to 700 live births1. They arise from disruptions in embryonic development influenced by genetic and environmental factors2. Beyond their aesthetic and functional impacts, orofacial clefts pose significant challenges in maintaining oral and systemic health, often leading to altered oral microbiota and increased vulnerability to infections3.
The oral microbiome is a highly diverse microbial community that plays a crucial role in maintaining oral and systemic health4. A stable oral microbiome contributes to preventing the colonization of pathogenic bacteria through competitive exclusion and the production of antimicrobial compounds. However, individuals with orofacial clefts experience disruptions in this balance, commonly referred to as dysbiosis. Factors such as surgical interventions, difficulties in maintaining oral hygiene due to structural abnormalities, feeding challenges, and frequent use of antibiotics contribute to these microbial changes5.
Among the microbial species that may dominate in dysbiotic states, Staphylococcus spp. are particularly noteworthy. These bacteria, including Staphylococcus aureus and coagulase-negative Staphylococci, are known for their ability to colonize mucosal surfaces and their role in various infections6. Staphylococcus aureus is a prominent human pathogen associated with a wide spectrum of clinical manifestations, including skin and soft tissue infections, bacteremia, endocarditis, pneumonia, and foodborne illnesses. In contrast, coagulase-negative Staphylococci are primarily implicated in infections related to prosthetic devices and indwelling medical implants7. The ability of these species to produce biofilms and exhibit antibiotic resistance further complicates their clinical management8.
Research suggests that children with orofacial clefts may have an increased prevalence of Staphylococcus spp. in their oral microbiomes9. This could be attributed to factors such as prolonged hospitalization, increased medical procedures, and compromised mucosal integrity10. To accurately identify and characterize these bacteria, advanced diagnostic techniques such as polymerase chain reaction (PCR) and Vitek assays are employed11. This study aimed to determine the interrater agreement between the molecular assay and the Vitek Compact assay for the detection of the prevalence of Staphylococcus aureus in Iraqi children suffering from orofacial clefts.
MATERIALS AND METHODS
Study area: This study was designed as a comparative cohort investigation conducted at Al-Qadisiya Teaching Hospital over the period of 9 months/September 2024 to May 2025. The study population comprised 200 pediatric patients diagnosed with orofacial clefts, specifically, cleft lip, cleft palate, or combined cleft lip and palate, confirmed through clinical examination by a qualified specialist or certified medical technician.
Participants included children aged 3 to 12 years who had not received systemic antibiotic therapy within the previous 30 days and who had no documented systemic illnesses. Exclusion criteria encompassed children presenting with active oral or systemic infections, additional congenital anomalies, or whose parents or guardians declined participation.
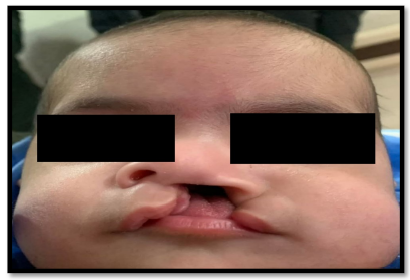
Sample collection: Oral swab samples were collected from each participant to target bacterial colonization in distinct oral sites via lip mucosa: Inner lip, often associated with general microbial load for cleft lip, and gingival crevices for cleft palate. Sterile cotton-tipped applicators were used for sample collection Fig. 1. For children with cleft palate, special care was taken to swab near the cleft site without causing discomfort or contamination. Samples were stored in sterile transport media at 4°C and transported promptly to the microbiology laboratory in the biology science college12.
|
The image demonstrates typical anatomical features of cleft lip and/or cleft palate used for clinical diagnosis and patient classification before oral sample collection in the present study
Bacterial isolation and identification: Sterile conditions were maintained for all equipment and procedures related to the handling of clinical samples. Each oral swab sample was directly inoculated onto Mannitol Salt Agar (MSA) and Blood Agar plates under aseptic conditions. The plates were incubated at 37°C for 24-48 hrs. Colony growth on primary cultures was evaluated based on morphological characteristics, including: Beta-hemolysis: Observed on Blood Agar as a clear zone around colonies, Mannitol fermentation: Identified on MSA by the production of yellow colonies due to acid production, as well as catalase test: All isolates were tested for catalase activity, confirming the presence of Staphylococcus spp. (catalase-positive), Coagulase test: Coagulase-positive isolates were presumptively identified as Staphylococcus aureus, and Gram staining: Gram-positive isolates displayed characteristic grape-like clusters under the microscope13.
Molecular study: For DNA extraction, subcultured the isolates onto Mueller-Hinton agar. After allowing for growth, we suspended five colonies in 100 μL of Tris-EDTA buffer and heated the mixture to 100°C. We then centrifuged the solution at 9000 xg for 30 sec, using 2 μL of the supernatant as the template in a 50 μL reaction. The primers used were mecA-F: 5’-GTAGAAATGACTGAACGTCCGATGA and mecA-R: 5’-CCAATTCCACATTGTTTCGGTCTAA14. The master mix included reaction buffer, MgCl2, dNTPs, mecA primers, Taq polymerase, and distilled water. We amplified the DNA using a thermocycler (Biobase, China) and subjected the amplicons to electrophoresis on a 1.5% agarose gel containing 0.5 μg/mL ethidium bromide. The gel was visualized under ultraviolet light, with positive tests showing a PCR product of 310 bp15.
Vitek compact2 analysis: Isolates of Staphylococcus aureus were obtained from culture plates and prepared for analysis with the Vitek Compact device in Al-Qadisiya Hospital. The results were compiled into a sheet and printed using Excel 201016.
Statistical analysis: Data were analyzed using SPSS software. Agreement between PCR and Vitek Compact 2 was assessed using Cohen’s kappa coefficient. Associations between cleft type and Staphylococcus positivity were evaluated using odds ratios (ORs) with 95% confidence intervals. A p<0.05 was considered statistically significant.
RESULTS AND DISCUSSION
In Table 1 The age groups were categorized as follows: 2-5, 6-8, and 9-12 years. A total of 60 participants, representing 30%, were included in the 2-5 years age group9-11. The 6-8-year age group was represented by 76 participants, accounting for 38% of the total. Sixty-four participants, corresponding to 32%, were classified in the 9-12 years age group17. In total, 200 participants were included in the study, representing 100% of the sample.
| Table 1: | Age-Wise distribution of participants | |||
| Age group (year) | Number of participants (n) | Percentage |
| 02-May | 60 | 30 |
| 06-Aug | 76 | 38 |
| 09-Dec | 64 | 32 |
| Total | 200 | 100 |
| Table 2: | Bacterial prevalence based on sampling areas | |||
| Sampling area | No. | Bacterial prevalence (n) | Percentage |
| Cleft palate | 100 | 39 | 60.9 |
| Cleft lip | 100 | 25 | 39.1 |
| Total | 200 | 64 | 100 |
| Table 3: | Distribution of Staphylococcus aureus positivity in soft and hard palate among cleft palate cases (n = 100) | |||
| Cleft palate | Positive for Staphylococcus (outcome) | Negative for Staphylococcus (outcome) | Total |
| Soft | 21 (True positives) | 16 (False negatives) | 37 |
| Hard | 18 (False positives) | 45 (True negatives) | 63 |
| Total | 39 (Total positive) | 61 (Total negative) | 100 |
| Table 4: | Staphylococcus aureus by cleft lip type | |||
| Cleft lip | Positive for Staphylococcus (outcome) | Negative for Staphylococcus (outcome) | Total |
| Unilateral | 13 (True positives) | 26 (False negatives) | 39 |
| Bilateral | 12 (False positives) | 49 (True negatives) | 61 |
| Total | 25 (Total positive) | 75 (Total negative) | 100 |
| The Kappa coefficient indicates strong agreement between the PCR and Vitek methods, reflecting a high proportion of true positives and true negatives with relatively few false positives and false negatives | |||
The sampling areas were divided into cleft palate and cleft lip, with each group consisting of 100 patients. In Table 2, A total of 39 positive results for Staphylococcus aureus were obtained from patients with cleft palate, representing 60.9% of the total. Additionally, 25 positive results were obtained from patients with cleft lip, accounting for 39.1% of the total18. In summary, 64 positive results were identified from the 200 patients sampled, representing 100% of the study population.
Table 2 presents data comparing the prevalence of Staphylococcus in children with soft and hard cleft palates. Among those with a soft cleft palate, a notable proportion tested positive for Staphylococcus, with fewer testing negative. In contrast, children with a hard cleft palate showed a smaller proportion of positive cases and a larger number of negative results. Overall, the data suggest a potential difference in the distribution of Staphylococcus positivity between the two groups, indicating a possible association between the type of cleft palate and the presence of Staphylococcus, odds ratio (OR) = 20.45: The odds of testing positive for Staphylococcus spp. are approximately 20.45 times higher for patients with Soft Cleft Palate compared to those with Hard Cleft Palate; This indicates that children with soft cleft palate have 3.28 times higher odds of being positive for Staphylococcus compared to those with hard cleft palate.
Table 3 shows that the soft palate had 21 positive and 16 negative cases, while the hard palate had 18 positive and 45 negative cases. Overall, S. aureus positivity was higher in the soft palate compared to the hard palate.
Table 4 presents the outcomes of Staphylococcus aureus testing in patients with cleft lip. Among patients with unilateral cleft lip, 13 true positive results were recorded for Staphylococcus aureus, alongside 26 false negatives, resulting in a total of 39 patients in this category. For those with bilateral cleft lip, 12 false positives were identified, while 49 true negatives were recorded, leading to a total of 61 patients. Overall, the study revealed 25 total positive results and 75 total negative results, summing to 100 patients assessed for the presence of Staphylococcus aureus. This suggests that cleft lip unilateral is a strong risk factor for testing positive for Staphylococcus aureus in a cohort of patients. OR = 2.02 means that the odds of testing positive for Staphylococcus spp. are about 2.02 times higher for patients with cleft lip unilateral compared to patients with cleft lip bilateral.
|
| Table 5: | Agreement between PCR and vitek assays | |||
| Rater agreement | PCR positive | PCR negative | Total |
| Vitek positive | 59 (True positives) | 29 (False positives) | 88 |
| Vitek negative | 5 (False negatives) | 130 (True negatives) | 112 |
| Total | 64 | 136 | 200 |
Unilateral cases had 13 positive and 26 negative results, while bilateral cases had 12 positive and 49 negative results. Positivity was higher in unilateral cleft lip, with strong agreement between methods (Kappa = 0.826).
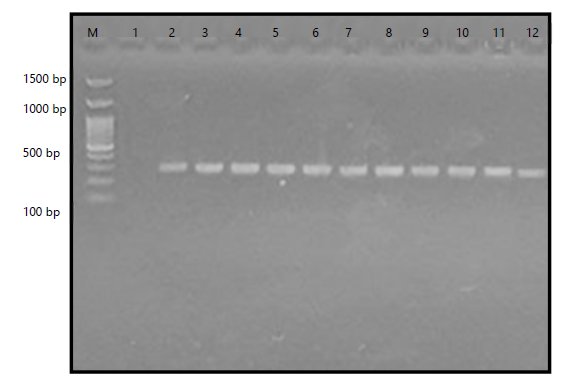
The gel electrophoresis results revealed distinct bands at 310 bp, confirming the presence of the mecA gene. A DNA ladder was used to estimate fragment sizes, ensuring accurate band identification. Positive samples displayed clear amplification, while negative controls showed no visible bands. Variations in band intensity were observed among samples, indicating differences in DNA concentration or amplification efficiency. No signs of contamination or non-specific amplification were detected. The separation of DNA fragments occurred as expected, with well-defined bands corresponding to the target mecA gene Fig. 2.
It has recently been discovered that polymicrobial colonies cause a disrupted and damaging response in the host through a worldwide process known as polymicrobial synergy and dysbiosis. Kids with CLP have an extremely diversified microbiome. Colonies of bacteria frequently work together to improve colonisation, perseverance, or virulence19.
The age distribution of participants in the study was categorized into three groups: 2-5, 6-8, and 9-12 years. A total of 200 participants were included, with the majority being in the 6-8 years age group (38%, n = 76). The 2-5 years age group was represented by 60 participants (30%), while the 9-12 years age group comprised 64 participants (32%). The distribution of participants across these age groups was considered to ensure a representative sample for the investigation of Staphylococcus aureus in orofacial clefts20. The data were analyzed to determine potential correlations between age and the prevalence of Staphylococcus aureus in children with orofacial clefts. It has been suggested that younger age groups, particularly those under 5 years, may exhibit higher susceptibility to bacterial colonization due to underdeveloped immune systems21. However, further investigation is required to confirm these findings, as the current study was limited by its sample size and demographic distribution.
The importance of age-specific analysis in microbiological studies has been emphasized in previous research, as age-related factors can significantly influence microbial colonization and infection rates22. The prevalence of Staphylococcus aureus was analyzed in two distinct sampling areas: Cleft palate and cleft lip. A total of 200 samples were collected, with 100 samples obtained from each area. The bacterial prevalence of Staphylococcus aureus was found to be higher in the cleft palate samples (60.9%, n = 39) compared to the cleft lip samples (39.1%, n = 25). These findings suggest that the cleft palate may provide a more favorable environment for bacterial colonization, potentially due to its anatomical complexity and increased susceptibility to microbial retention23.
The observed differences in bacterial prevalence between the two sampling areas were consistent with previous studies, which have reported that the cleft palate is more prone to infections due to its proximity to the nasal cavity and oral microbiota. The higher percentage of Staphylococcus aureus in cleft palate samples could also be attributed to the challenges in maintaining oral hygiene in this region, particularly in pediatric populations24.
The total prevalence of Staphylococcus aureus across both sampling areas was 32% (n = 64), indicating that bacterial colonization is a significant concern in individuals with orofacial clefts. These findings highlight the need for targeted interventions, such as improved hygiene practices and prophylactic measures, to reduce the risk of infections in this population25.
The presence of Staphylococcus aureus among individuals with cleft palate has been investigated, revealing significant findings regarding its occurrence in both soft and hard cleft palates. The data suggest that S. aureus was more frequently detected in cases involving soft cleft palates, with 21 individuals testing positive. Meanwhile, 16 individuals with soft cleft palates tested negative, indicating a notable proportion of cases where S. aureus was absent. In contrast, individuals with hard cleft palates exhibited a lower prevalence of S. aureus positivity, as only 18 individuals tested positive, whereas 45 tested negative. These findings suggest a possible association between the type of cleft palate and the likelihood of S. aureus colonization.
The overall data indicate that out of 100 individuals, 39 tested positive for S. aureus, while 61 tested negative. This distribution highlights the potential role of cleft palate morphology in bacterial colonization. The observed variation in bacterial presence between soft and hard cleft palates may be attributed to anatomical and physiological differences, which could influence microbial adherence and growth26. The soft palate, being more flexible and dynamic in function, may provide a more suitable environment for bacterial retention compared to the rigid structure of the hard palate.
Previous studies have suggested that individuals with cleft palates are more susceptible to bacterial colonization due to structural abnormalities, which may lead to increased bacterial reservoirs in oral and nasal cavities. The increased prevalence of S. aureus in soft cleft palates aligns with findings that demonstrate higher bacterial loads in regions with greater tissue mobility. Additionally, variations in hygiene practices, surgical interventions, and antibiotic use could contribute to differences in bacterial colonization rates27.
The total data for cleft lip indicate that out of 100 individuals, 25 tested positive for S. aureus, while 75 tested negative. The variation in colonization rates between unilateral and bilateral cleft lip cases may be influenced by structural and functional differences affecting bacterial adherence. Factors such as hygiene practices, surgical interventions, and antibiotic use could further contribute to differences in bacterial colonization rates20.
The agreement between PCR and Vitek assay results for the detection of S. aureus has also been assessed using the Kappa coefficient. The findings indicate that 59 cases were classified as true positives by both PCR and Vitek, whereas 29 cases were identified as false positives by Vitek. Additionally, 5 cases were classified as false negatives by Vitek, while 130 cases were considered true negatives. The total number of cases analyzed was 200. The overall agreement between the two assays provides insight into the reliability of Vitek as a diagnostic tool in comparison to PCR28.
The Kappa coefficient is frequently used to measure inter-rater agreement, and in this study, it reflects the degree of consistency between the two methods. A strong agreement suggests that Vitek could serve as a viable alternative to PCR in clinical settings where rapid results are necessary. However, the presence of false positives and false negatives indicates that discrepancies exist between the two methodologies. These variations may arise due to differences in detection sensitivity, specificity, or external factors such as sample quality and contamination risks. Further investigation is warranted to refine diagnostic accuracy and establish optimal protocols for bacterial identification in clinical practice29.
The detection of the mecA gene by PCR has been widely utilized to confirm methicillin resistance in Staphylococcus aureus. In the gel electrophoresis image, bands at 310 bp were observed, indicating the successful amplification of the mecA gene in multiple samples. The presence of these bands suggests that the bacterial isolates tested carry the mecA gene, which encodes penicillin-binding protein 2a (PBP2a) and confers resistance to β-lactam antibiotics, including methicillin. The identification of mecA-positive isolates is essential for guiding antibiotic therapy and implementing infection control measures in clinical settings30.
The PCR has been recognized as a highly sensitive and specific method for detecting antibiotic resistance genes. The interpretation of the gel image relies on the comparison of the sample bands with a DNA ladder, which serves as a molecular weight reference. In this case, the bands detected at 310 bp confirm the presence of the mecA gene in the tested isolates. If negative controls were included in the experiment, their absence of bands would validate the reliability of the results and exclude the possibility of contamination. However, false negatives may occur due to PCR inhibitors, insufficient DNA extraction, or sequence variations affecting primer binding sites31.
The clinical significance of mecA detection lies in its role in identifying methicillin-resistant S. aureus (MRSA), a major cause of hospital- and community-acquired infections. The rapid and accurate identification of mecA-positive isolates is critical for selecting appropriate antimicrobial therapy aence of S. aureus among individuals with cleft lind reducing the spread of resistant strains. In addition to PCR, other methods such as PBP2a latex agglutination assays and whole-genome sequencing have been employed for further validation. Despite its high sensitivity, PCR may occasionally yield false positives due to non-specific amplification, necessitating the use of confirmatory tests32.
CONCLUSION
This study demonstrates a notable prevalence of Staphylococcus aureus colonization among Iraqi children with orofacial clefts, with a higher occurrence in cleft palate cases compared to cleft lip. The detection of the mecA gene confirms the presence of methicillin-resistant strains, indicating a potential challenge for clinical management and antibiotic therapy. The strong agreement between PCR and the Vitek system supports the reliability of both molecular and automated methods for accurate bacterial identification. Overall, the findings emphasize the need for improved infection control, prudent antibiotic use, and routine microbial surveillance in cleft care, while future research should assess the long-term effects of surgical and preventive interventions on oral microbial profiles.
SIGNIFICANCE STATEMENT
This study underscores the clinical importance of detecting methicillin-resistant Staphylococcus aureus in patients with cleft lip and palate, revealing a notable prevalence of mecA-positive isolates. Combining molecular PCR and automated Vitek diagnostics enhances accuracy, enabling timely identification of antibiotic resistance, guiding effective antimicrobial therapy, and supporting infection control measures in this vulnerable population, ultimately reducing the risk of resistant infections.
ACKNOWLEDGMENT
The authors acknowledged their gratitude to the College of Science, University of Al-Qadisiya, Iraq.
REFERENCES
- Mossey, P.A., J. Little, R.G. Munger, M.J. Dixon and W.C. Shaw, 2009. Cleft lip and palate. Lancet, 374: 1773-1785.
- Dixon, M.J., M.L. Marazita, T.H. Beaty and J.C. Murray, 2011. Cleft lip and palate: Understanding genetic and environmental influences. Nat. Rev. Genet., 12: 167-178.
- Kilian, M., I.L.C. Chapple, M. Hannig, P.D. Marsh and V. Meuric et al., 2016. The oral microbiome-an update for oral healthcare professionals. Br. Dent. J., 221: 657-666.
- Dewhirst, F.E., T. Chen, J. Izard, B.J. Paster and A.C.R. Tanner et al., 2010. The human oral microbiome. J. Bacteriol., 192: 5002-5017.
- Otto, M., 2008. Staphylococcal biofilms. Curr. Topics Microbiol. Immun., 322: 207-228.
- Tong, S.Y., J.S. Davis, E. Eichenberger, T.L. Holland and V.G. Fowler Jr, 2015. Staphylococcus aureus infections: Epidemiology, pathophysiology, clinical manifestations, and management. Clin. Microbiol. Rev., 28: 603-661.
- Lowy, F.D., 1998. Staphylococcus aureus infections. N. Engl. J. Med., 339: 520-532.
- Archer, N.K., M.J. Mazaitis, J.W. Costerton, J.G. Leid, M.E. Powers and M.E. Shirtliff, 2011. Staphylococcus aureus biofilms: Properties, regulation and roles in human disease. Virulence, 2: 445-459.
- Świtała, J., M. Sycińska-Dziarnowska, G. Spagnuolo, K. Woźniak, K. Mańkowska and L. Szyszka-Sommerfeld, 2023. Oral microbiota in children with cleft lip and palate: A systematic review. J. Clin. Med., 12.
- Chandrasekaran, S., S.S. Rao and V. Shetty, 2025. Association between oral microbiome in children undergoing palatoplasty and surgical site infection. Cleft Palate Craniofacial J.
- Adesina, O.A., A.A. Efunkoya, K.U. Omeje and P.I. Idon, 2016. Postoperative complications from primary repair of cleft lip and palate in a semi-urban Nigerian teaching hospital. Niger. Med. J., 57: 155-159.
- Katayama, Y., T. Ito and K. Hiramatsu, 2002. A new class of genetic element, staphylococcus cassette chromosome mec, encodes methicillin resistance in Staphylococcus aureus. Antimicrob. Agents Chemother., 44: 1549-1555.
- Chambers, H.F. and F.R. deLeo, 2009. Waves of resistance: Staphylococcus aureus in the antibiotic era. Nat. Rev. Microbiol., 7: 629-641.
- Pantosti, A., 2012. Methicillin-resistant Staphylococcus aureus associated with animals and its relevance to human health. Front. Microbiol., 3.
- Martins, K.B., A.M. Ferreira, A.L. Mondelli, T.T. Rocchetti and M. de L.R. de S da Cunha, 2018. Evaluation of MALDI-TOF VITEK®MS and VITEK® 2 system for the identification of Staphylococcus saprophyticus. Future Microbiol., 13: 1603-1609.
- Lee, J., E. Baek, H. Ahn, J. Bae and S. Kim et al., 2024. Development of a one-step multiplex qPCR assay for detection of methicillin and vancomycin drug resistance genes in antibiotic-resistant bacteria. Pathogens, 13.
- Jiang, W., Z. Yan, Z. Chen, L. Gu and H. Bao et al., 2024. Investigating oral microbiome profiles in patients with cleft lip and palate compared with the healthy control. BMC Oral Health, 24.
- Escobar-Arregocés, F., M.A. Eras, A. Bustos, A. Suárez-Castillo, D.A. García-Robayo and M. del Pilar Bernal, 2024. Characterization of the oral microbiota and the relationship of the oral microbiota with the dental and periodontal status in children and adolescents with nonsyndromic cleft lip and palate. Systematic literature review and meta-analysis. Clin. Oral Invest., 28.
- Simon, A.K., G.A. Hollander and A. McMichael, 2015. Evolution of the immune system in humans from infancy to old age. Proc. Biol. Sci., 282.
- Nasidze, I., J. Li, D. Quinque, K. Tang and M. Stoneking, 2009. Global diversity in the human salivary microbiome. Genome Res., 19: 636-643.
- Seidel, C.L., K. Strobel, M. Weider, M. Tschaftari and C. Unertl et al., 2023. Orofacial clefts alter early life oral microbiome maturation towards higher levels of potentially pathogenic species: A prospective observational study. J. Oral Microbiol., 15.
- Landis, R.J. and G.G. Koch, 1977. The measurement of observer agreement for categorical data. Biometrics, 33: 159-174.
- Oliveira, D.C. and H. de Lencastre, 2002. Multiplex PCR strategy for rapid identification of structural types and variants of the mec element in methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother., 46: 2155-2161.
- Chen, W., J. Zhang, H. Wei, J. Su and J. Lin et al., 2024. Rapid and sensitive detection of methicillin-resistant Staphylococcus aureus through the RPA-PfAgo system. Front. Microbiol., 15.
- Brook, I., 2008. Microbiology and management of soft tissue and muscle infections. Int. J. Surg., 6: 328-338.
- Wertheim, H.F.L., D.C. Melles, M.C. Vos, W. van Leeuwen, A. van Belkum, H.A. Verbrugh and J.L. Nouwen, 2005. The role of nasal carriage in Staphylococcus aureus infections. Lancet Infect. Dis., 5: 751-762.
- Peacock, S.J. and G.K. Paterson, 2015. Mechanisms of methicillin resistance in Staphylococcus aureus. Annu. Rev. Biochem., 84: 577-601.
- Kalan, L. and E.A. Grice, 2018. Fungi in the wound microbiome. Adv. Wound Care, 7: 247-255.
- Zaura, E., E.A. Nicu, B.P. Krom and B.J.F. Keijser, 2014. Acquiring and maintaining a normal oral microbiome: Current perspective. Front. Cell. Infect. Microbiol., 4.
- The Human Microbiome Project Consortium, 2012. Structure, function and diversity of the healthy human microbiome. Nature, 486: 207-214.
- Gao, L., T. Xu, G. Huang, S. Jiang, Y. Gu and F. Chen, 2018. Oral microbiomes: More and more importance in oral cavity and whole body. Protein Cell, 9: 488-500.
- Krismer, B., C. Weidenmaier, A. Zipperer and A. Peschel, 2017. The commensal lifestyle of Staphylococcus aureus and its interactions with the nasal microbiota. Nat. Rev. Microbiol., 15: 675-687.
How to Cite this paper?
APA-7 Style
Hussein Abass,
H.A., Kareem Ataia,
Z.A., Muhammed,
H.A. (2026). Prevalence of Staphylococcus aureus in the Oral Microbiome of Children with Orofacial Clefts: A Comparative Study using PCR and Vitek Assays. Asian Journal of Biological Sciences, 19(2), 70-78. https://doi.org/10.3923/ajbs.2026.70.78
ACS Style
Hussein Abass,
H.A.; Kareem Ataia,
Z.A.; Muhammed,
H.A. Prevalence of Staphylococcus aureus in the Oral Microbiome of Children with Orofacial Clefts: A Comparative Study using PCR and Vitek Assays. Asian J. Biol. Sci 2026, 19, 70-78. https://doi.org/10.3923/ajbs.2026.70.78
AMA Style
Hussein Abass
HA, Kareem Ataia
ZA, Muhammed
HA. Prevalence of Staphylococcus aureus in the Oral Microbiome of Children with Orofacial Clefts: A Comparative Study using PCR and Vitek Assays. Asian Journal of Biological Sciences. 2026; 19(2): 70-78. https://doi.org/10.3923/ajbs.2026.70.78
Chicago/Turabian Style
Hussein Abass, Hayder, Abdul, Zainab Abdul Kareem Ataia, and Hayder Ali Muhammed.
2026. "Prevalence of Staphylococcus aureus in the Oral Microbiome of Children with Orofacial Clefts: A Comparative Study using PCR and Vitek Assays" Asian Journal of Biological Sciences 19, no. 2: 70-78. https://doi.org/10.3923/ajbs.2026.70.78

This work is licensed under a Creative Commons Attribution 4.0 International License.


